Research
The Center for Disease Neurogenomics (CDN, director: Panos Roussos) has an established track record of successful collaborations and numerous joint publications. Our work within CDN integrates single-cell and spatial transcriptomics to uncover cell-type-specific regulatory mechanisms in disease states across multiple disease-focused consortia, including PsychENCODE, AMP-AD, AMP-PD, PsychAD, BD2, and TargetALS. We primarily focus on developing and applying computational and machine learning methodologies for understanding the genetic basis of neuropsychiatric and neurodegenerative disorders.
Single Cell Neurogenomics
Single-cell data is expanding in terms of depth and breadth; the current technology allows simultaneous measurement of multi-omic profiles from the same cell across a large breadth of samples. As a result, data analysis is becoming computationally demanding and requires more sophisticated methods such as machine learning to capture the underlying biological and pathological processes at the cellular level. The team will oversee the analysis of single-cell data, including but not limited to transcriptomic, regulatory, and proteomic profiles generated at the center. The team will build expertise in developing and adopting computationally efficient and effective algorithms to better understand the etiology and progression of neuropsychiatric and neurodegenerative disorders.
PsychAD
We play a key role in advancing large-scale, multi-omics studies to identify molecular pathways underlying disease risk and progression. One example is the PsychAD Project (R01AG067025). To gain deeper insights into the etiology of AD and related neurodegenerative disorders, we generated one of the largest single-cell transcriptomic atlas of the human dorsolateral prefrontal cortex (DLPFC) from PsychAD cohort (in press at Nature), profiled from 1,494 donors, spanning eight major neurodegenerative and neuropsychiatric diseases. This population-scale cross-disorder dataset was the foundational resource for characterizing the transcriptomic, genetic, and cellular vulnerability of AD by comparing it against other brain disorders. In particular, we focused on identifying the shared and distinct patterns of transcriptomic perturbations and the genetic factors that lead to co-pathologies. Characterizing shared vulnerabilities and pathophysiology has significant implications for early treatment and the development of effective therapeutics. Transcriptomic variation across multiple diseases pointed toward a central process involving the neuro-immune-vascular systems. One key insight was that the brain vascular system is intricately linked to immune dysfunction and that vascular cells were a key mediator of neurodegenerative processes in AD.
Machine Learning
We frame biological problems using mathematical and statistical tools. Leveraging the wealth of single-nucleus and functional genomic data, we apply interdisciplinary approaches like statistical modeling and machine learning to interpret disease genomes.
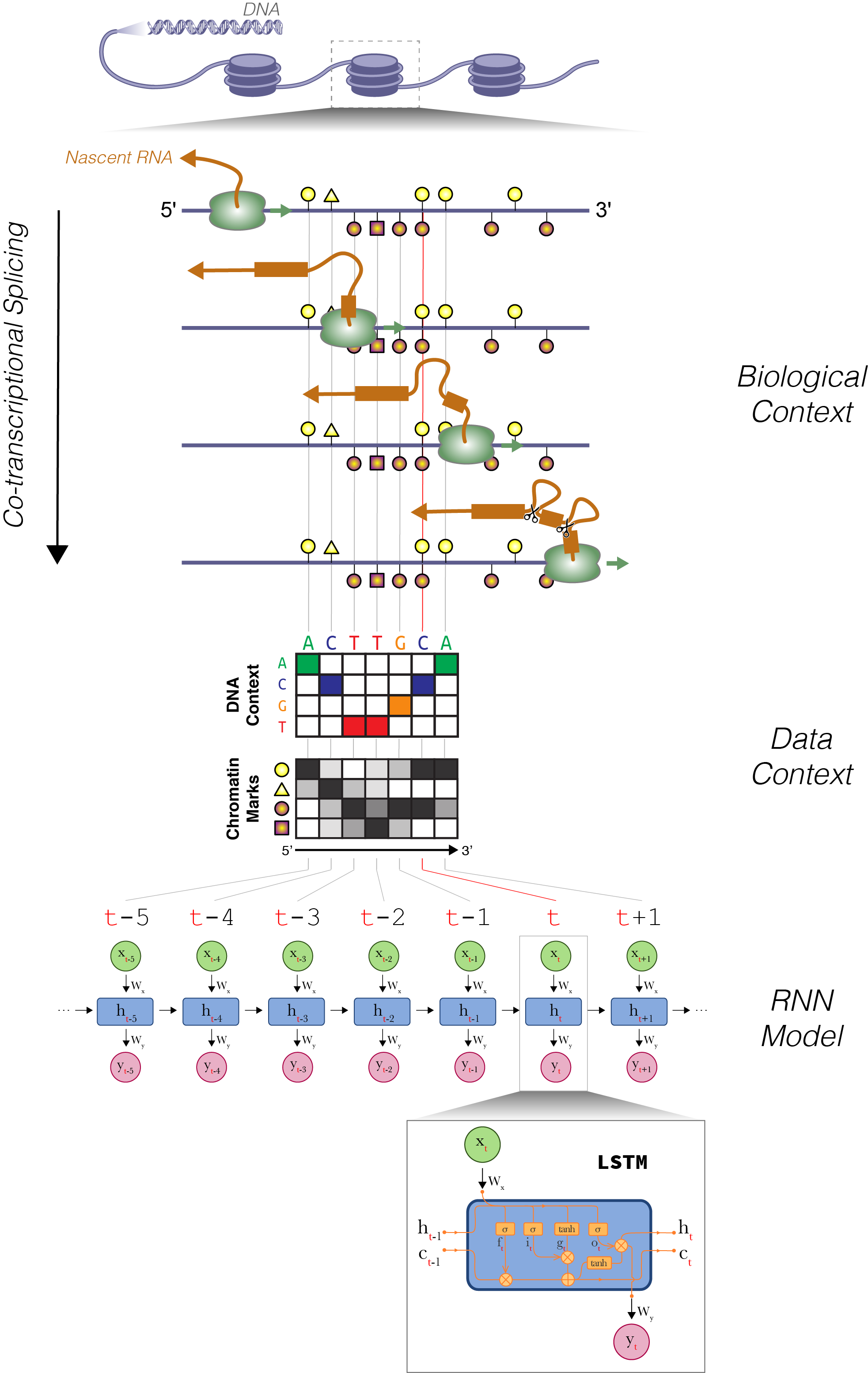
Regulatory Networks and Dynamics
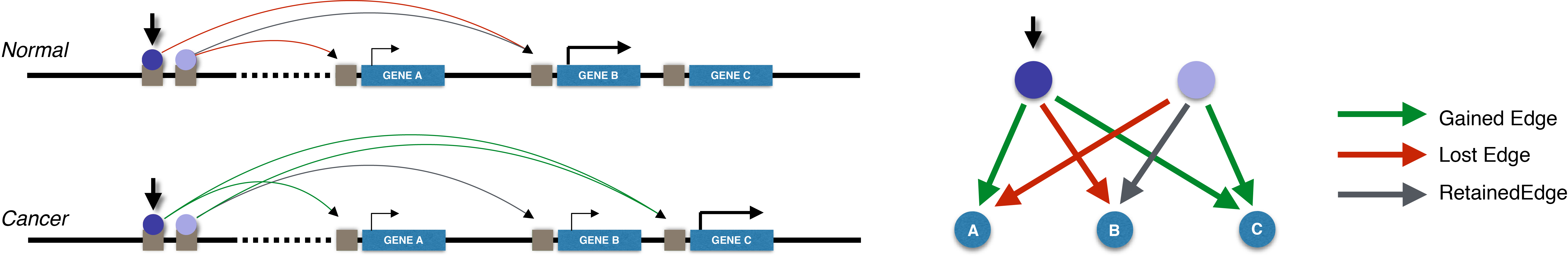
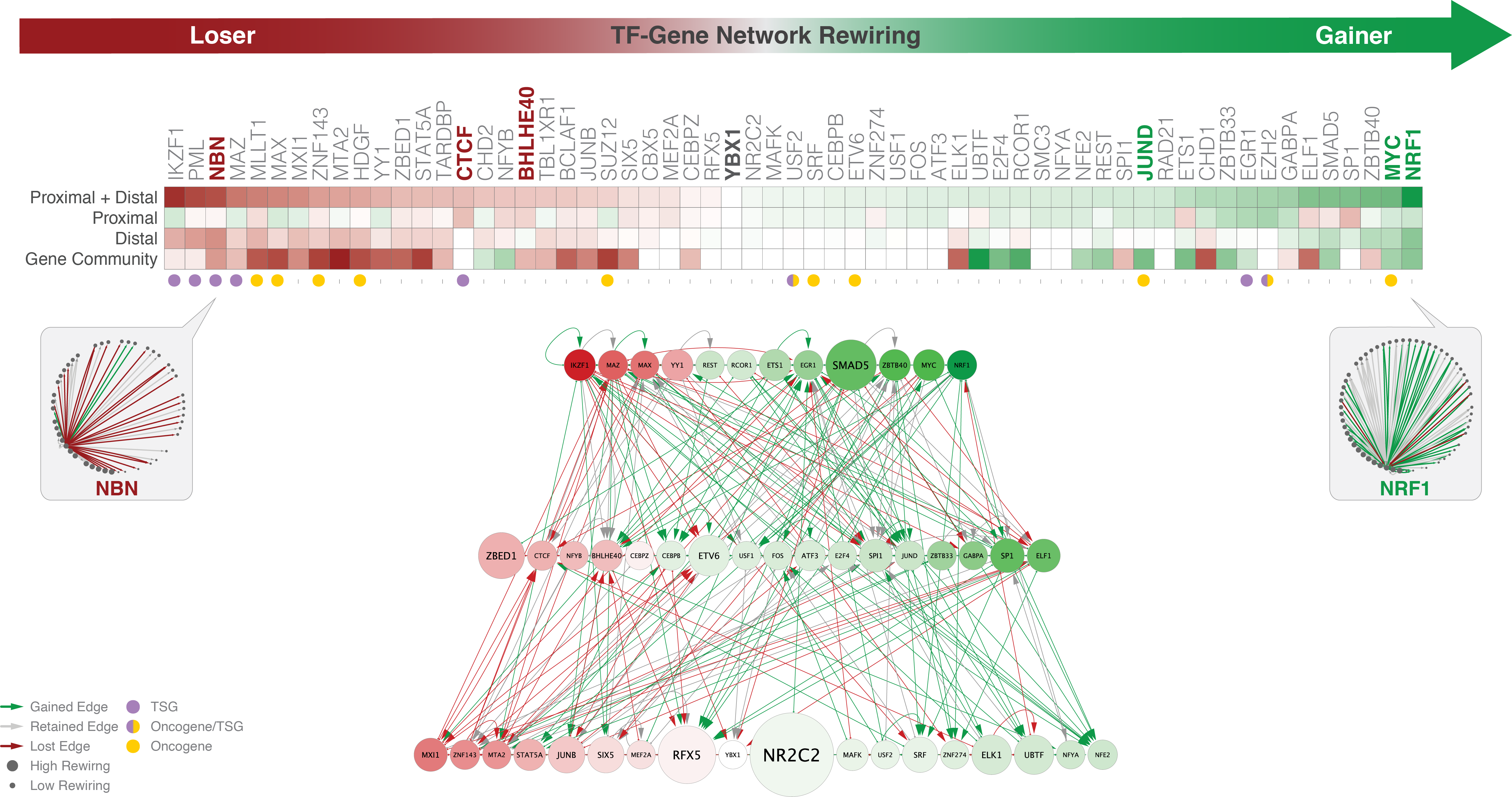
Integrative approaches can uncover meaningful patterns and rules from large data. We identify cis-regulatory elements, build gene regulatory networks, and discover regulatory dynamics during transcription.